Studies in hamsters are limited by the relative unavailability of tools to conduct immunological studies. We expect that these libraries will be a valuable resource and provide additional useful microsatellite markers for both the coral host and zooxanthellae.īackground The Syrian hamster, Mesocricetus auratus, has distinct immunological features and is uniquely susceptible to intracellular pathogens. Expected heterozygosities ranged from 0.38-0.85 and 0.08-0.87 for A. Using 25 individuals per species collected from each French Frigate Shoals (FF) and Johnston Atoll (JO), the number of alleles per locus ranged from 2-9. cytherea, and 8 microsatellites and the atpsβ locus from M. 
We report testing and optimization for 7 of these microsatellites from A. In addition, we also present redesigned primers for the nuclear coding region (atpsβ) for use in M. Blast searches indicate that these libraries contain microsatellites from both the coral host and Symbiodinium endosymbionts from each coral. Here we report the development of microsatellite marker libraries from holobiont extracts of four corals: Acropora cytherea (n = 50), Fungia scutaria (n = 118), Montipora capitata (n = 140) and Porites lobata (n = 149). The Hawaiian archipelago offers a unique perspective into understanding the population genetic structuring of this ecologically important group of organisms. Emerg Infect dis 4(3), 382–389.Comprehensive population genetic studies of coral communities are comparatively rare because of the scarcity of population genetic markers for many coral species. (1998) Detection and identification of previously unrecognized microbial pathogens. (2004) Identification of regions in multiple sequence alignments thermodynamically suitable for targeting by consensus oligonucleotides: application to HIV genome. 
(1999) Multiplex detection of single-nucleotide variations using molecular beacons. 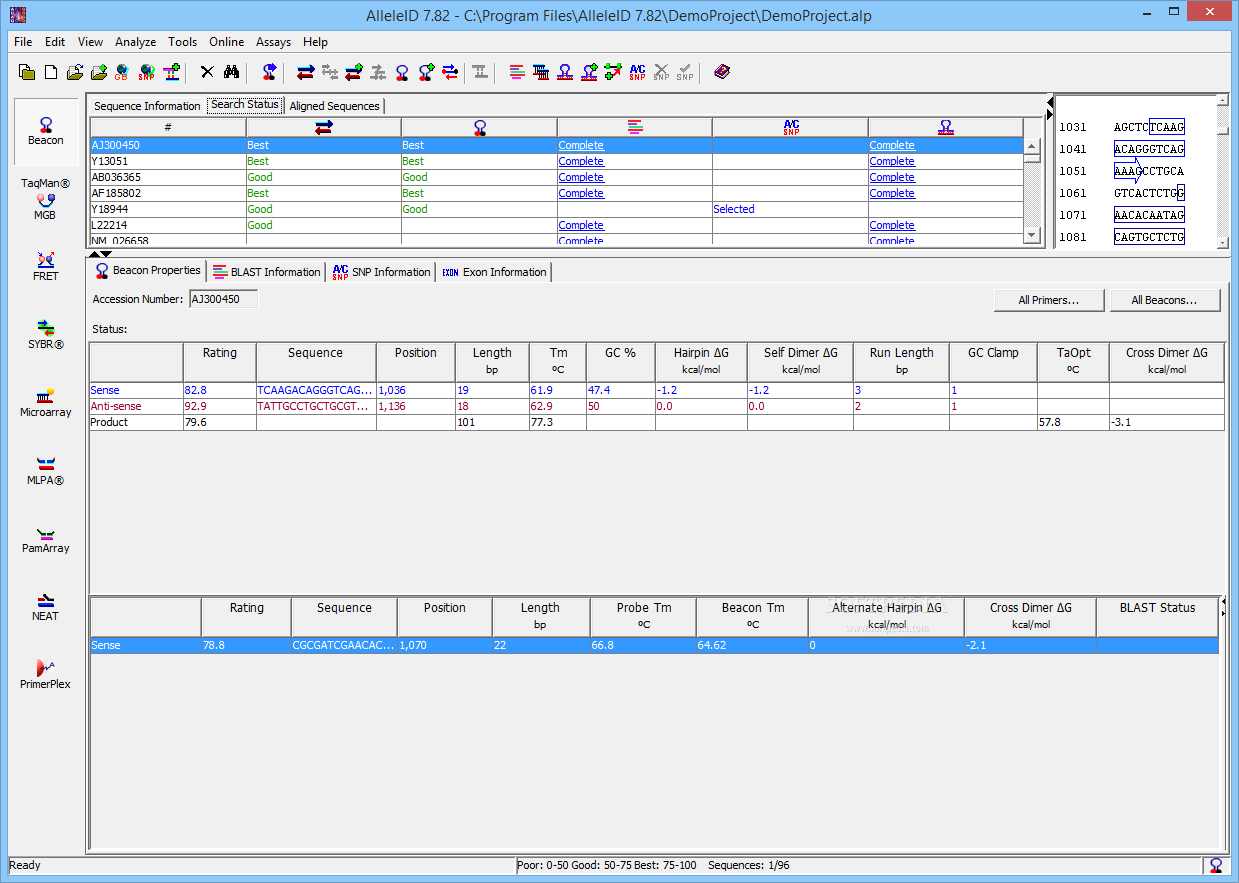
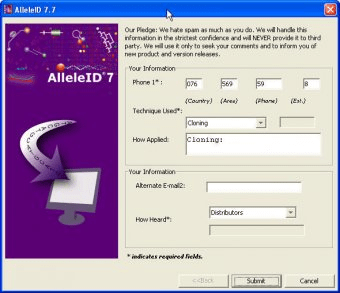
(1999) Multiplex detection of four pathogenic retroviruses using molecular beacons. R., Marras, S.A., Tyagi, S., Dube, S., Poiesz, B.J., et al. (2004) Simultaneous detection of Entamoeba histolytica, Giardia lamblia, and Cryptosporidium parvum in fecal samples by using multiplex real-time PCR. A., Templeton, K., Schinkel, J., Brienen E. (1991) Detection of specific polymerase chain reaction product by utilizing 5 ′-3 ′ exonuclease activity of Thermus aquaticus DNA polymerase. (1998) CLUSTAL: a package for performing multiple sequence alignment on a microcomputer. (2005) Quantitative detection of Listeria monocytogenes in biofilms by real-time PCR. Guilbaud, M., Coppet, P., Bourion, F., Rachman, C. (2005) Diagnostic system for rapid and sensitive differential detection of pathogens. Key Wordsīriese, P., Palacios, G., Kokoris, M., Jabado, O., Liu, Z., Renwick, N., Kapoor, V., Casas, I., Pozo, F., Limberger, R. Here, we describe a program AlleleID that designs qPCR and microarray assays to identify and detect pathogens. Manual design of these primer and probes is both tedious and results in lower quality of results because of the inability to simultaneously handle multiple criteria for design. Both of these techniques have a high accuracy when used with specific primers and probes. Although numerous chemistries are available for the molecular level identification of pathogens, the most common are qPCR and DNA microarrays. Sequence-based molecular methods such as real-time PCR, microarrays, and band biosensors provide high sensitivity, rapid diagnostics, and higher specificity allowing differentiation between related strains. Nucleic acid-based methods for the detection of microorganisms are rapid, sensitive and are generally successful even when the culturing of microorganisms fails. Conventional methods have longer turnaround time and, in most cases, lower sensitivity. Efficient clinical diagnosis of pathogens is important for the management of infectious diseases.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
